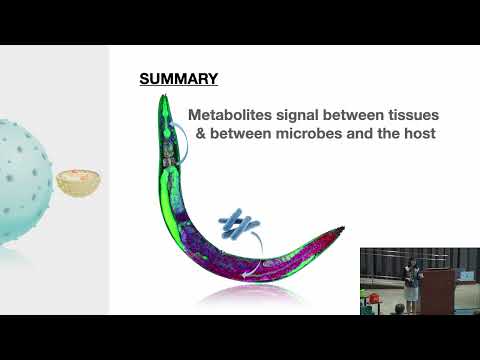
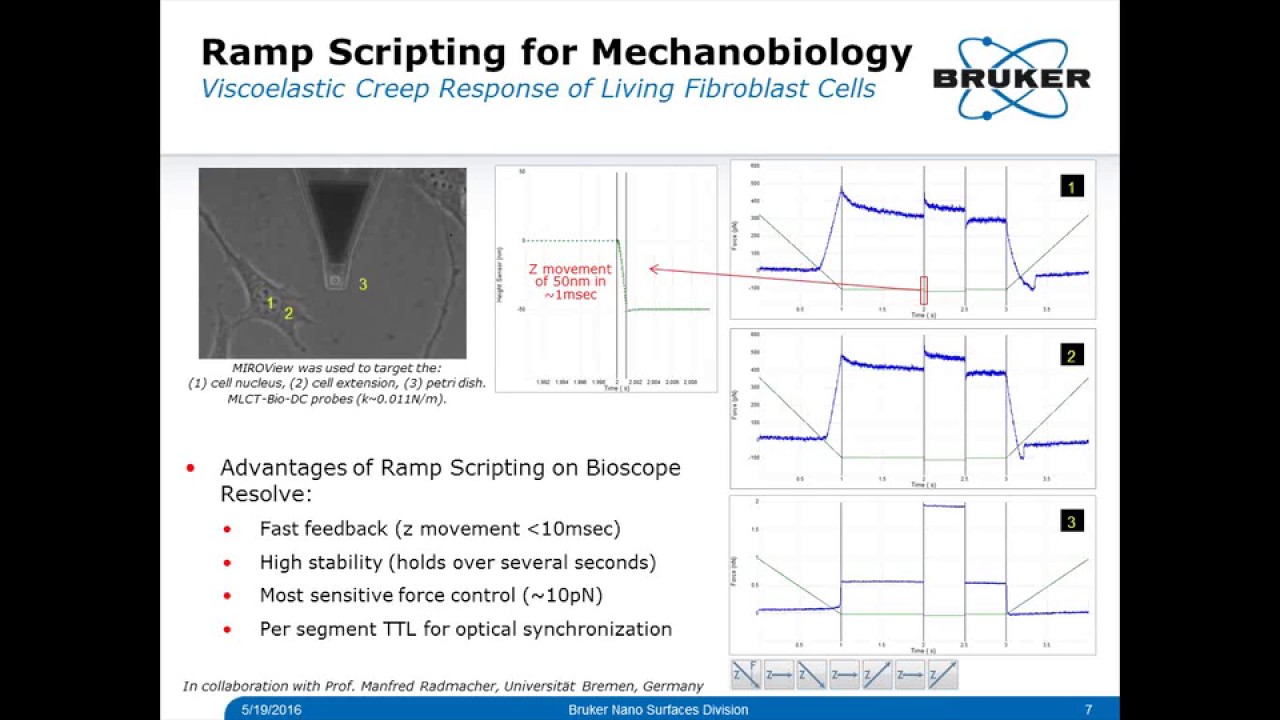
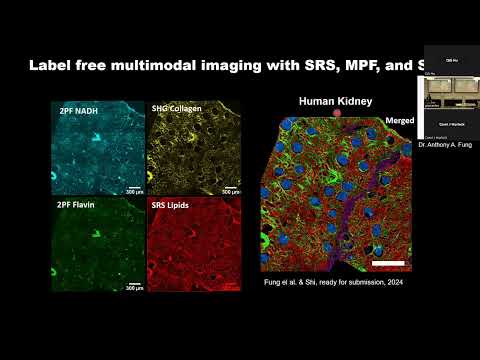
I think that you made a very important point Alex, that many researchers may not even have access to advanced imaging technology but it seems that it would be so useful to revisit and observe what effect specific therapies have at a cellular level.
Also, it looks like you referenced quite a few people who are doing research this way. I think it will be important to have people who can summarize the current research and draw conclusions because I think often times researchers may become hyperfocused or biased to their own specific niche.
Another question that I have been considering a lot lately is how presbyopia and cataracts form. These are age-related eye conditions which happen to everyone eventually. Traditionally it is taught that the lens in the eye grows like an onion and the cells in the middle are old, the lens then gets bigger, less flexible and cloudy. I don’t think this is the whole story because cataract progression is highly variable in the population and apparently at least somewhat modifiable by many substances. So long story short, I came across this paper and it describes the discrepancies between different measurement techniques and also between in vitro and in vivo findings.
(https://www.ncbi.nlm.nih.gov/pmc/articles/PMC2871961/#__ffn_sectitle).
Also to the point about re-visiting the research, this paper is 10 years old and I couldn’t find anything recent with a better answer.
I think this is an important question because if the lens does indeed keep growing through life, like an onion, then it is going to be more difficult restore the youthful phenotype, but if it somehow maintains itself until damage accumulates then there is a potential to activate repair pathways. Also I think if the lens keeps growing, then treatment that is directed specifically to the lens will be needed, meaning that any single systemic therapy will be unlikely to fully restore the entire organism. Advances in vivo imaging could help find the answer to this important question.